| Alias |
SPL6 |
| TF Classification |
| FioreDB | SBP (Click to show phylogenetic tree) |
| RARTF | SBP |
| AtTFDB | SBP |
| PlnTFDB | SBP (v3.0), SBP (v1.0) |
| DATF | SBP |
|
| TAIR short description |
Squamosa promoter-binding protein-like (SBP domain) transcription factor family protein (.1) |
| TAIR annotation |
Squamosa promoter-binding protein-like (SBP domain) transcription factor family protein; FUNCTIONS IN: DNA binding, sequence-specific DNA binding transcription factor activity; INVOLVED IN: regulation of transcription; LOCATED IN: nucleus; EXPRESSED IN: 14 plant structures; EXPRESSED DURING: 6 growth stages; CONTAINS InterPro DOMAIN/s: Transcription factor, SBP-box (InterPro:IPR004333); BEST Arabidopsis thaliana protein match is: squamosa promoter binding protein-like 14 (TAIR:AT1G20980.1); Has 836 Blast hits to 835 proteins in 40 species: Archae - 0; Bacteria - 0; Metazoa - 2; Fungi - 0; Plants - 834; Viruses - 0; Other Eukaryotes - 0 (source: NCBI BLink). (.1) |
| Gene model |
|
| Entry clone (w/o stop) |
IE_04D06_x |
| External link |
|
| Gene Ontology (GO) |
| (P) 10468: other metabolic processes |
->regulation of gene expression |
| (F) 3677: DNA or RNA binding |
->DNA binding |
| (F) 3700: transcription factor activity |
->transcription factor activity, sequence-specific DNA binding |
| (P) 42742: response to abiotic or biotic stimulus |
->defense response to bacterium |
| (F) 46872: other binding |
->metal ion binding |
| (F) 5515: protein binding |
->protein binding |
| (C) 5634: nucleus |
->nucleus |
| (P) 6351: transcription,DNA-dependent |
->transcription, DNA-templated |
| (P) 6355: other metabolic processes |
->regulation of transcription, DNA-templated |
|
| miRNA/tasiRNA |
| Target sequence |
miRNA/tasiRNA sequence |
Map |
5'--3'
|
| miR156a: 3'-CACGAGTGAGAGAAGACAGT-5' |
|
 |
5'--3'
|
| miR156b: 3'-CACGAGTGAGAGAAGACAGT-5' |
|
 |
5'--3'
|
| miR156c: 3'-CACGAGTGAGAGAAGACAGT-5' |
|
 |
5'--3'
|
| miR156d: 3'-CACGAGTGAGAGAAGACAGT-5' |
|
 |
5'--3'
|
| miR156e: 3'-CACGAGTGAGAGAAGACAGT-5' |
|
 |
5'--3'
|
| miR156f: 3'-CACGAGTGAGAGAAGACAGT-5' |
|
 |
5'--3'
|
| miR156g: 3'-CACGAGTGAGAGAAGACAGC-5' |
|
 |
5'--3'
|
| miR156h: 3'-CACGAGAGAAAGAAGACAGTT-5' |
|
 |
5'--3'
|
| miR156i: 3'-CACGAGAGAGAGAAGACAGTT-5' |
|
 |
5'--3'
|
| miR156j: 3'-CACGAGAGAGAGAAGACAGT-5' |
|
 |
5'--3'
|
| miR157a: 3'-CACGAGAGATAGAAGACAGTT-5' |
|
 |
5'--3'
|
| miR157b: 3'-CACGAGAGATAGAAGACAGTT-5' |
|
 |
5'--3'
|
| miR157c: 3'-CACGAGAGATAGAAGACAGTT-5' |
|
 |
5'--3'
|
| miR157d: 3'-CACGAGAGATAGAAGACAGT-5' |
|
 |
|
| Orthologs |
|
Type |
| Ia |
Ib |
II |
III |
Total |
| poplar |
|
1 |
|
|
1 |
| soybean |
|
1 |
|
|
1 |
| rice |
|
|
|
|
|
| sorghum |
|
|
|
|
|
| physcomitrella |
|
|
|
|
|
| Total |
|
2 |
|
|
2 |
|
| Repression motif |
|
Repression motif
in putative orthologs |
|
| Status |
| Plasmid construction | -> done |
| Plasmid construction | -> done |
| Transformed | -> done |
| T1 seed harvested | -> done |
| T2 seed harvested | -> done |
|
CRES-T phenotype from individual project
View all photo |
no visible phenotype (direct input)
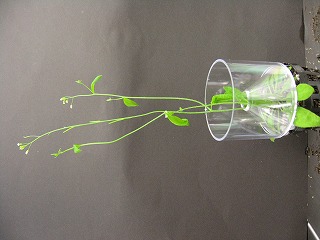      and more... and more... |
| CRES-T phenotype from publication |
|
| CRES-T phenotype from bulk project |
|
Phenotype in ornamental plants
|
|
| Comment |
|
